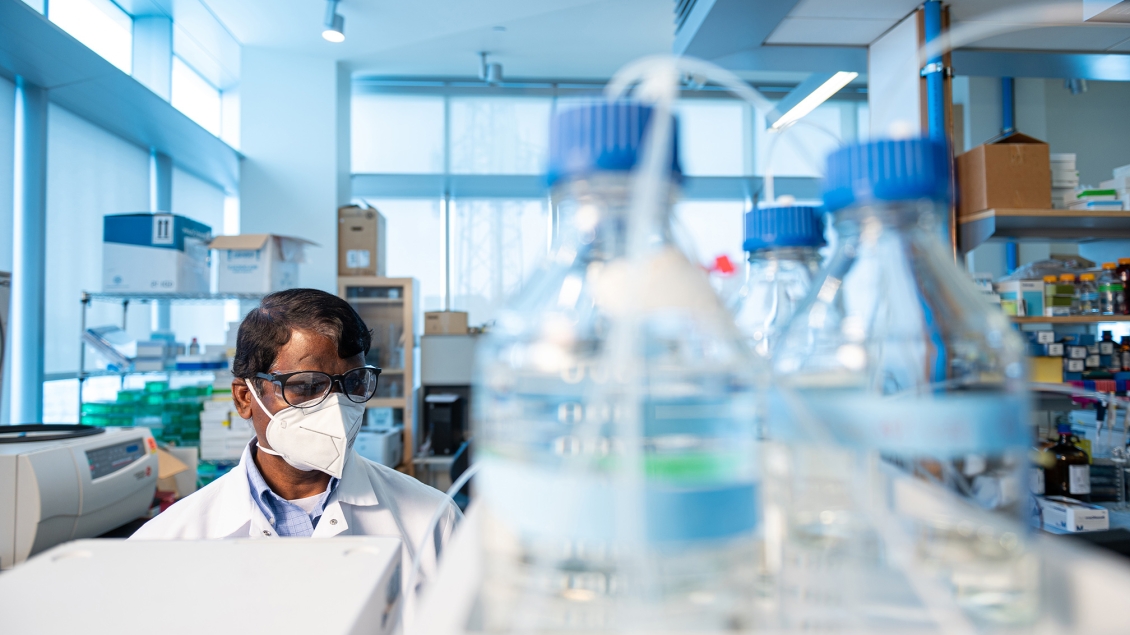
We measure concentrations of small molecules in biological samples on a recharge basis.
The Metabolomics Core strives to provide GLP-quality analytical services at competitive prices and regularly customize assays to fit our client’s needs. Services include initial consultation, method development, data collection, interpretation, and presentation. Core competencies include both targeted assays for many analyte classes as well as untargeted assays to evaluate the entire metabolome in biofluids and tissues, measurement of flux through various pathways using 13C-isotopomer analyses, and structural identification of isolated unknowns and potential biomarkers.
The Metabolomics Core, funded by the NIH Common Fund’s Metabolomics program, is a fully integrated center that supports metabolomics research and training nationwide.
We work with other Regional Comprehensive Metabolomics Resource Cores (RCMRC)s and the Metabolomics Consortium Data Repository and Coordinating Center (DRCC) to develop standard protocols for metabolomic data annotation, storage, and dissemination.
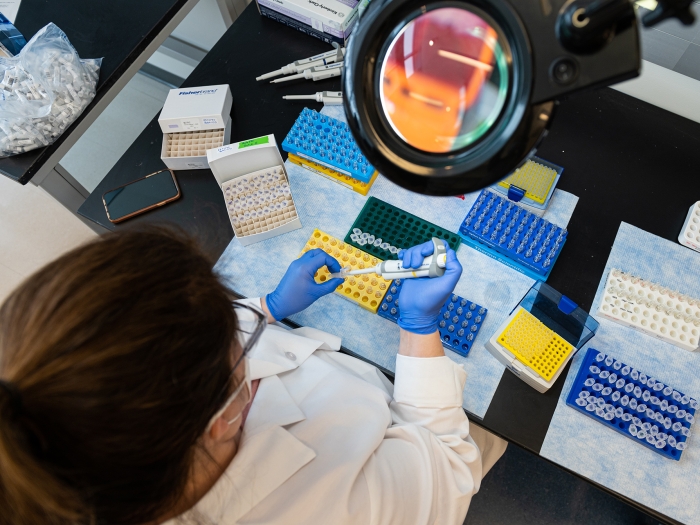

Services include initial consultation, method development, data collection, interpretation, and presentation.
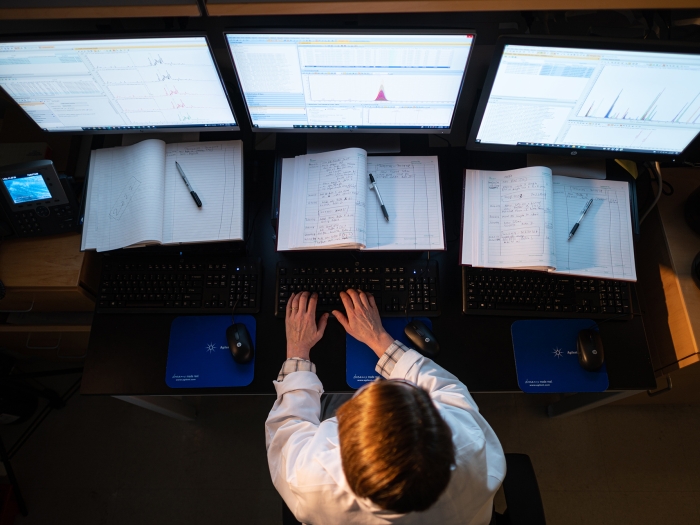

We provide a variety of educational opportunities for students, fellows, and faculty in the field of metabolomics research, as well as offer access to various funding mechanisms.
Quotes for all projects are based on a standard list of prices and assay descriptions found below. Analytes important to your project can be added to any standard assay, and new assays can be developed to suit your requirements. Please contact Managing Director Maureen Kachman to discuss the details.
*A surcharge will be applied to all external non-U-M customers.
| Service | 1-50 Samples | 51-100 Samples | 101+ Samples | Assay Description |
| 2-HG - D/L isomer analysis | $151 | $121 | $109 | Quantification of the D/L isomers of 2-Hydroxglutarate after derivatization (which allows for separation on LC-MS.). All analytes and Internal Standards are measured on a LC-QQQ using MRM methods. Analytes reported as uM and normalized to either wet tissue weight or cell protein content. |
| Acylcarnitine analysis | $144 | $129 | $116 | Profile 30 acylcarnitine species in plasma, serum, or tissues after solvent extraction. All analytes and Internal Standards are measured on a LC-QQQ using MRM methods. Analytes reported as uM and normalized to either wet tissue weight or cell protein content. |
| Amino acid analysis (free amino acids) | $144 | $129 | $116 | Profile 20 free amino acids in plasma, serum, cells, tissues, or feces after solvent extraction. All analytes and Internal Standards are measured on a LC-QQQ using MRM methods. Analytes reported as uM and normalized to either wet tissue weight or cell protein content. |
| Amino acid analysis (with protein hydrolysis) | $187 | $150 | $135 | Profile 20 amino acid species in plasma, serum, or tissues after solvent extraction, including protein hydrolysis to determine the entire amino acid contents of a sample. All analytes and Internal Standards are measured on an LC-QQQ using MRM methods. Analytes reported as uM and normalized to either wet tissue weight or cell protein content. |
| Bile acid analysis | $151 | $135 | $122 | Profile 24 bile acid species in plasma, serum, tissues, or feces after solvent extraction. All analytes and Internal Standards are measured on an LC-QQQ using MRM methods. Analytes reported as uM and normalized to either wet tissue/fecal weight. |
| Glycolysis / TCA / Nucleotide analysis | $167 | $144 | $130 | Profile TCA (citate, malate, succinate, AKG, fumarate) and glycolysis (glucose, G/F6P, FBP, 2/3PG, PEP, and pyruvate) and selected nucleotide and energy metabolites in tissue, cells, or plasma (AKG, fumarate, pyruvate, and phosphometabolites may be undetectable in plasma.) All analytes and Internal Standards are measured on an accurate mass QTOF. Analytes reported as uM and normalized to either wet tissue/feces weight or cell protein content. |
| Eicosanoids (Oxylipins) | $158 | $141 | $123 | Profile 28 Eicosanoids in tissue, including HETEs, HODEs, LTKB4, TBXB2, and Prostaglandins. Eicosanoids are extracted and concentrated using solid phase extraction and measured on an LC-QQQ using MRM methods. Analytes reported as uM and normalized to either wet tissue weight or cell protein content. |
| Tryptophan Metabolites analysis | $158 | $126 | $114 | Profile 6 Tryptophan pathway metabolites in plasma, serum, tissues, or feces after solvent extraction. Hydroxy species may be difficult to measure in plasma and cells. All analytes and Internal Standards are measured on an LC-QQQ using MRM methods. Analytes reported as uM and normalized to either wet tissue/feces weight or cell protein content. |
| Short chain fatty acid analysis | $137 | $122 | $107 | Profile 6 Short Chain Fatty acids in plasma, serum, tissues, or feces after solvent extraction. Great care must be taken to isolate samples from lab sources of very low MW fatty acids, especially for plasma. Please consult the Core before beginning sample collection. All analytes and Internal Standards are measured on an LC-QQQ using MRM methods. Analytes reported as uM and normalized to either wet tissue/feces weight or cell protein content. |
| Steroid Hormone Panel - Delta4 | $191 | $173 | $155 | Profile 24 Delta-4 steroids, including testosterone(s), estrogen(s), cortisone/sol, and progesterone(s) in biofluids (cells/solid tissue analysis not available at this time). Please consult the Core if estrogen measurements are required for normal individuals, a high-sensitivity method must be employed. All analytes and Internal Standards are measured on an LC-QQQ using MRM methods, with analytes reported as uM values. |
| 13C mass isotopologue (flux) analysis - TCA | $167 | $134 | $120 | Isotope enrichment measurements on the metabolites reported in the Gly/TCA/Nucleotides platform above., plus key amino acids. Isotope enrichment of targeted metabolites are corrected for natural abundance and reported as ratios of M, M+1, M+2, M+3, etc. Please consult the Core on the choice of label (if not U13C Glucose) or any other questions (e.g., labeling times or substate amounts) before submitting samples for analysis. |
| 13C mass isotopologue (flux) analysis - Acylcarnitine | $167 | $134 | $120 | Isotope enrichment measurements on the metabolites reported in the Acylcarnitine platform above. Isotope enrichment of targeted metabolites are corrected for natural abundance and reported as ratios of M, M+1, M+2, M+3, etc. Please consult the Core on the choice of label (usually U13C oleate or U13C palmitate) or any other questions (e.g., labeling times or substrate amounts) before submitting samples for analysis. |
| 13C mass isotopologue (flux) analysis - Other | $167 | $144 | $121 | Isotope enrichment measurements on the metabolites reported in any other established platform. Please consult the Core on the choice of label or any other questions (e.g., labeling times or substrate amounts) before submitting samples for analysis. |
| Custom Panel One | $127 | $114 | $101 | Measure up to 3 custom metabolites in one LC-MS run (one mode/one form of chromatography). Additional charges may be necessary for method development or the purchase of project-specific isotope-labeled internal standards. |
| Custom Panel Two | $144 | $129 | $116 | Measure up to 6 custom metabolites in two LCMS runs (two modes or two forms of chromatography. Additional charges may be necessary for method development or the purchase of project-specific isotope-labeled internal standards. |
| Custom Panel Three | $179 | $143 | $129 | Measure up to 10 custom metabolites in two LCMS runs (two modes or two forms of chromatography). Additional charges may be necessary for method development or the purchase of project-specific isotope-labeled internal standards. |
| Service | 1-50 Samples | 51-100 Samples | 101+ Samples | Assay Description |
| Untargeted Metabolomics - "Broad Coverage" (biofluids) | $329 | $283 | $221 | Untargeted Metabolite Profiling using Reversed Phase Chromatography / QTOF in Positive and Negative modes. Approximately 350 named metabolites reported include Amino acids, Acylcarnitines, Fatty acids, Lipids, Bile acids, etc. Analytes measured are reported as peak area per sample for both known (annotated) compounds and unknown features. Basic statistical analysis is included. Unknown identification is done on statistically significant features and is aided by the acquisition of comprehensive MS/MS. |
| Untargeted Metabolomics - "Broad Coverage" (tissue) | $359 | $309 | $241 | See description for Untargeted Metabolomics (biofluids). Tissues and other solid samples are normalized by accurately weighing samples and scaling the extraction solvent volume to the measured weight. For cells, an accurate cell count is requested for normalization. |
| Central Carbon Metabolism - "Polar Coverage" (biofluids) | $142 | $123 | $98 | Untargeted Metabolite Profiling using Ion Pairing Chromatography / QTOF in Negative mode. Approximately 80 named metabolites reported include compounds in the Amino acids, Glycolysis, and TCA pathways, as well as Nucleotides and other Organic acids. Analytes measured are reported as peak area per sample for both known (annotated) compounds and unknown features. Basic statistical analysis is included. Unknown identification is done on statistically significant features and is aided by the acquisition of comprehensive MS/MS. |
| Central Carbon Metabolism - "Polar Coverage" (tissue) | $174 | $150 | $118 | See description for Central Carbon Metabolism Profiling (biofluids). Tissues and other solid samples are normalized by accurately weighing samples and scaling the extraction solvent volume to the measured weight. For cells, an accurate cell count is requested for normalization. |
| Untargeted Broad + Polar Coverage (biofluids) | $375 | $339 | $278 | Untargeted Metabolite Profiling using both Reversed Phase and Ion Pairing Chromatography / QTOF with Positive and Negative modes (2). Approximately 400 named metabolites reported include Amino acids, Acylcarnitines, Fatty acids, Lipids, Bile acids, Glycolysis and TCA pathways, Nucleotides, Organic acids, and others. Analytes measured are reported as peak area per sample for both known (annotated) compounds and unknown features. Basic statistical analysis is included. Unknown identification is done on statistically significant features and is aided by the acquisition of comprehensive MS/MS. |
| Untargeted Broad + Polar Coverage (tissue) | $388 | $367 | $300 | See description for Central Carbon Metabolism Profiling (biofluids.) Tissues and other solid samples are normalized by accurately weighing samples and scaling the extraction solvent volume to the measured weight. For cells, an accurate cell count is requested for normalization. |
| Service | 1-50 Samples | 51-100 Samples | 101+ Samples | Assay Description |
| Shotgun Lipidomic (biofluids) | $349 | $312 | $276 | Nontargeted Lipid Profiling using Reversed Phase Chromatography in Positive and Negative modes. Typically >800 lipids are reported from ~30 lipid classes. Lipids are reported with their lipid class, total chain length, and total double bond count. A basic data analysis report with unsupervised visualization and differential analysis is included. Targeted lipidomics is available for specific individual lipid species with method development and additional costs. For complex study designs, we recommend investigators engage a statistician or consult with the Core if our statistician’s services are desired. |
| Shotgun Lipidomic (tissue) | $390 | $348 | $313 | See description for Shotgun Lipidomics (biofluids). For tissues and other solid samples, data are normalized to accurate tissue weights (recorded at the Core). For cells, an accurate cell count is requested for normalization. |
| Service | Rate |
| Data Analysis (hourly) | $44/per hour |
We Offer Access to State-of-the-Art Technology
- Multiple high-resolution, high mass accuracy qTOF
- High sensitivity triple-quad instruments for LC/MS
- Mass unit resolution instruments for GC/MS
- QQQ boasts a triple quadrupole performance known industry-wide for meeting quantitative requirements with sensitivity, renowned reliability, and overall system robustness
- A laboratory information management system (MetLIMS) that captures study metadata, study samples, sample processing history, and final results
- A bioinformatics and mathematical modeling infrastructure that visualizes and models complex omics-level interactions with the phenotype
How to Initiate a Project
The process for initiating a project with the Metabolomics Core varies depending on your institutional affiliation. In the appropriate section below, please review the guidelines and forms. Once the necessary steps are completed, your request will be reviewed. A consultation will be scheduled to review the project/experimental design with you and to provide a quote.
Refer to our rates for assay descriptions and standard pricing.
For general guidelines in sample collection for metabolomics studies, and specific instructions on how to submit your samples, please download these important documents:
NOTE: For metabolomics experiments with smaller sample sets (<20) for common assays, there may be a delay in analysis. In order to maintain overall laboratory efficiency, small sets will be added to larger sample runs or pooled with other small sample sets for analysis. Core users may inquire about expedited analysis of smaller numbers of samples.
To request services, users must log in to MiCores. If you don’t already have an account, register for one here.
MiCores (iLabs software, part of Agilent Technologies) is an online system used by the U-M Medical School’s Biomedical Research Core Facilities (BRCF) to streamline the process of ordering and billing for core service requests. For instructions on setting up a MiCores account and tutorials on the system, visit the MiCores Learning Site.
Once you have a MiCores account, you must request access to the Metabolomics Core facility. Your institutional affiliation (i.e., 3a, 3b, or 3c below) will then determine how to proceed.
For general MiCores questions:
Call the HITS Service desk: (734) 936-8000
Select prompt #3
Select prompt #3 again
For password resets:
Call the HITS Service desk: (734) 936-8000
Select prompt #3
Select prompt #1
Or visit the HITS website for more assistance.
Metabolomics Project Request (U-M Affiliated)
Investigators affiliated with the University of Michigan will be charged the advertised rate for all services, and do not need to sign a contract. A U-M shortcode for billing is required when initiating any analyses.
Metabolomics Project Request (Non-U-M Investigator)
Investigators not affiliated with the University of Michigan will need to complete the MC Service Agreement below. After an initial consultation, an ‘Exhibit A’ will be drafted by the Core that specifies the scope and total costs for your project, and is part of the MC Service agreement. A copy of U-M’s W9 form is also found below. These 3 documents (Exhibit A, Service Agreement, and W-9) should be forwarded to the appropriate authority at your institution. A PO number and copies of the signed contract must be e-mailed to the Core Administrator before services can be initiated. A surcharge will be applied to all external non-U-M customers. Please contact the Managing Director, Maureen Kachman, for more information.
Further information can be found in the documents listed below:
Services for All For-profit Institutions
A more elaborate contract is required for all for-profit clients. We can start with your standard contract, and negotiate some of the terms and conditions, if necessary with the U-M Office of Contract Administration.
Please contact the Managing Director, Maureen Kachman, to discuss details.



1000 Wall Street
Ann Arbor, MI 48109
The Metabolomics Core is one of the Biomedical Research Core Facilities, and a part of the Medical School Office of Research, where our mission is to foster an environment of innovation and efficiency that serves the Michigan Medicine research community and supports biomedical science from insight to impact.